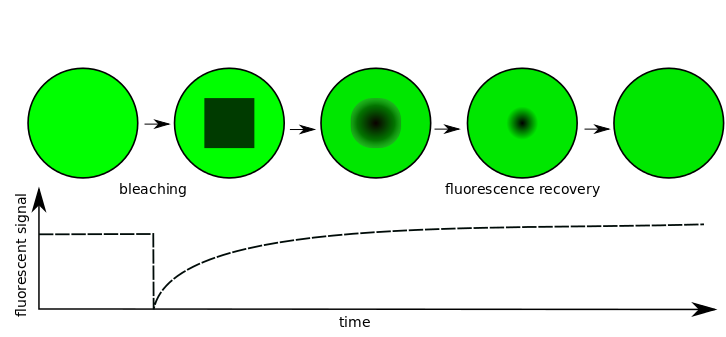
Fluorescence Recovery After Photobleaching (FRAP) is a common technique to assess molecular diffusion. PyFRAP is a novel simulation-based analysis software that makes use of PDE simulations to analyze FRAP experiments in 3D geometries. It uses the first post-bleach image as initial condition, making assumptions about the underlying initial conditions obsolete. PyFRAP can fit different reaction-diffusion models to FRAP data, providing quantitative information about effective diffusion.
PyFRAP comes with a full image analysis and simulation toolbox. In particular, PyFRAP can
- Import FRAP datasets from timelapse experiments and analyze image data with various options such as
- image filters
- background subtraction
- illumination correction
- Simulate the FRAP experiment with exact interpolated initial conditions
- Fit simulated experiment to analyzed data and extract diffusion coefficient
- Statistical analysis of fitting results
- Hierarchical data structure making data exchange/sharing easy
- Comprehensive GUI, making almost all PyFRAP tools available
This is the most recent developmental branch of PyFRAP (Python 3). It offers two options:
- PyFRAP migrated to Python 3 (tested in Ubuntu 20.04, Windows 10/11, macOS 10.15).
- A portable Linux/Ubuntu-based virtual machine, which is sufficiently isolated from forced (breaking) Python updates. This option also provides pre-installed accessory programs like gmsh, which is helpful for developing new geometries.
PyFRAP can be installed in different ways. Installation instructions can be found here.
If you are familiar with Python and git, you can install PyFRAP via
git clone https://github.com/mueller-lab/PyFRAP
git checkout crystal
and:
python setup.py install --user
We highly recommend installing with the --user option, since PyFRAP needs to read/write data files in the installation folder. In some cases, this might lead to file permission issues.
To have a tested/robust installation on all major platforms, please head over to SysBiolWorkbench.readme and carefully follow the instructions.
PyFRAP depends on
- pyopenssl
- ndg-httpsclient
- pyasn1
- ez_setup>=0.9
- numpy>=1.23.2
- scipy>=0.13.3
- matplotlib>=1.4.3
- scikit-image>=0.11.3
- FiPy>=3.1
- PyQt5
- colorama>=0.2.5
- VTK>=9.2.0rc1
- numpy-stl
- solidpython
- wget
- python-bioformats
PyFRAP comes with a comprehensive GUI. It can easily be started by clicking on runPyFRAP.bat (Windows), runPyFRAP.command (OSX) or runPyFRAP.sh (Linux).
If you are using a terminal, cd to your PyFRAP directory and type
python pyfrp/PyFRAP.py
If you are already in a Python session, you can simply run
import pyfrp
pyfrp.main()
Note that in the latter method PyFRAP's stdout might get redirected to the Python shell you are executing it from.
PyFRAP is a complete Python package and can be imported via
import pyfrpNote that PyFRAP has three main submodules: pyfrp.modules, pyfrp.subclasses and pyfrp.gui. If you do not want to use any GUI elements, we recommend only importing the modules you need.
Check out the PyFRAP wiki's First Steps Section.
PyFRAP is fully documented, allowing easy creation of scripts and extensions of the PyFRAP toolbox. The API of PyFRAP can be found here .
To learn more about PyFRAP, check out the PyFRAP Wiki.